import numpy as np
import torch
import torch.nn as nn
from scipy.integrate import odeint as scipy_odeint
import matplotlib.pyplot as plt
# Reproducibility
torch.manual_seed(42)
np.random.seed(42)
plt.rcParams.update({
"figure.dpi": 130,
"axes.spines.top": False,
"axes.spines.right": False,
"axes.grid": True,
"grid.alpha": 0.3,
"font.size": 8,
"axes.titlesize": 8,
"axes.labelsize": 8,
"xtick.labelsize": 7,
"ytick.labelsize": 7,
"legend.fontsize": 7,
})
C_S, C_I, C_R = "#2196F3", "#F44336", "#4CAF50"
C_OBS, C_PINN = "black", "#9C27B0"
# --- Ground truth parameters ---
N = 1_000
beta_true = 0.3
gamma_true = 0.1
T_max = 60.0
y0 = np.array([990.0, 10.0, 0.0])
def sir_rhs(y, t, beta, gamma, N):
S, I, R = y
return [-beta*S*I/N, beta*S*I/N - gamma*I, gamma*I]
n_grid = 100
t_grid = np.linspace(0, T_max, n_grid)
sol = scipy_odeint(sir_rhs, y0, t_grid, args=(beta_true, gamma_true, N))
S_true, I_true, R_true = sol.T
print(f"R₀ = {beta_true/gamma_true:.1f}")
print(f"Peak infections: {I_true.max():.0f} on day {t_grid[I_true.argmax()]:.1f}")
print(f"Final susceptibles: {S_true[-1]:.0f}")
# --- Sparse, noisy observations of I(t) only ---
n_obs = 30
obs_idx = np.sort(np.random.choice(np.arange(1, n_grid), n_obs, replace=False))
t_obs = t_grid[obs_idx]
I_obs = np.clip(I_true[obs_idx] + np.random.normal(0, 20.0, n_obs), 0, N)
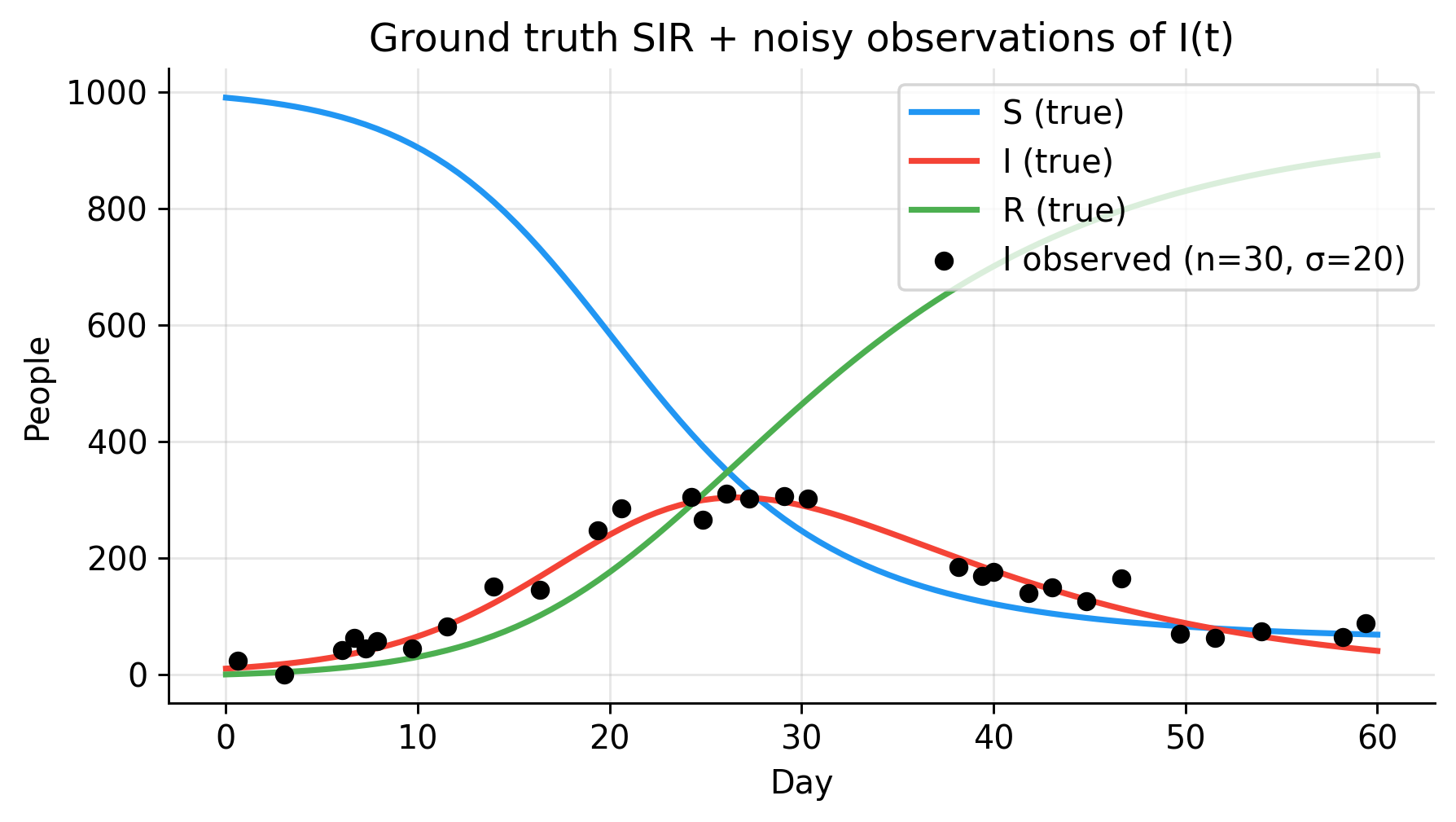
fig, ax = plt.subplots(figsize=(5, 3))
ax.plot(t_grid, S_true, color=C_S, lw=2, label="S (true)")
ax.plot(t_grid, I_true, color=C_I, lw=2, label="I (true)")
ax.plot(t_grid, R_true, color=C_R, lw=2, label="R (true)")
ax.scatter(t_obs, I_obs, color=C_OBS, s=30, zorder=5,
label=f"I observed (n={n_obs}, σ=20)")
ax.set(xlabel="Day", ylabel="People",
title="Ground truth SIR + noisy observations of I(t)")
ax.legend(loc="upper right")
plt.tight_layout()
plt.show()