# ── True SIR for Neural ODE / PINN extrapolation experiment ──────────────────
beta_t = 0.3
gamma_t = 0.1
T_max = 60.0
T_train = 40.0
I0_frac = 1e-3
def sir_rhs_np(y, t):
s, i, r = y
return [-beta_t*s*i, beta_t*s*i - gamma_t*i, gamma_t*i]
y0_sir = [1 - I0_frac, I0_frac, 0.0]
t_full = np.linspace(0, T_max, 400)
sol_sir = scipy_odeint(sir_rhs_np, y0_sir, t_full)
# 25 noisy I observations in [0, T_train]
n_obs = 25
t_tr = t_full[t_full < T_train]
obs_idx = np.sort(np.random.choice(len(t_tr) - 1, n_obs, replace=False))
t_obs_np = t_tr[obs_idx]
i_noisy = np.clip(
sol_sir[t_full < T_train][obs_idx, 1]
+ np.random.normal(0, 0.012, n_obs), 0, 1)
# ── SEIR with NPI for UDE experiment ─────────────────────────────────────────
sigma_t = 1 / 5.0
BETA_HIGH2, BETA_LOW2, NPI_DAY2 = 0.35, 0.14, 25.0
T_seir = 70.0
T_seir_tr = 45.0
def seir_rhs_np(y, t):
s, e, i, r = y
b = BETA_HIGH2 if t < NPI_DAY2 else BETA_LOW2
return [-b*s*i, b*s*i - sigma_t*e, sigma_t*e - gamma_t*i, gamma_t*i]
y0_seir = [1 - I0_frac, 0.0, I0_frac, 0.0]
t_seir = np.linspace(0, T_seir, 500)
sol_seir = scipy_odeint(seir_rhs_np, y0_seir, t_seir)
n_obs2 = 25
t_seir_tr_arr = t_seir[t_seir < T_seir_tr]
obs_idx2 = np.sort(
np.random.choice(len(t_seir_tr_arr) - 1, n_obs2, replace=False))
t_obs2_np = t_seir_tr_arr[obs_idx2]
i_noisy2 = np.clip(
sol_seir[t_seir < T_seir_tr][obs_idx2, 2]
+ np.random.normal(0, 0.012, n_obs2), 0, 1)
# ── Tensors ───────────────────────────────────────────────────────────────────
N_STEPS = 50 # Euler integration steps (fast; increase for higher accuracy)
t_eval = torch.linspace(0, T_train, N_STEPS)
t_eval_f = torch.linspace(0, T_max, 400)
t_obs_t = torch.tensor(t_obs_np, dtype=torch.float32)
i_obs_t = torch.tensor(i_noisy, dtype=torch.float32)
obs_t_idx = [int(round(float(t) / T_train * (N_STEPS - 1))) for t in t_obs_np]
t_eval2 = torch.linspace(0, T_seir_tr, N_STEPS)
t_eval2_f = torch.linspace(0, T_seir, 500)
t_obs2_t = torch.tensor(t_obs2_np, dtype=torch.float32)
i_obs2_t = torch.tensor(i_noisy2, dtype=torch.float32)
obs2_t_idx = [int(round(float(t) / T_seir_tr * (N_STEPS - 1))) for t in t_obs2_np]
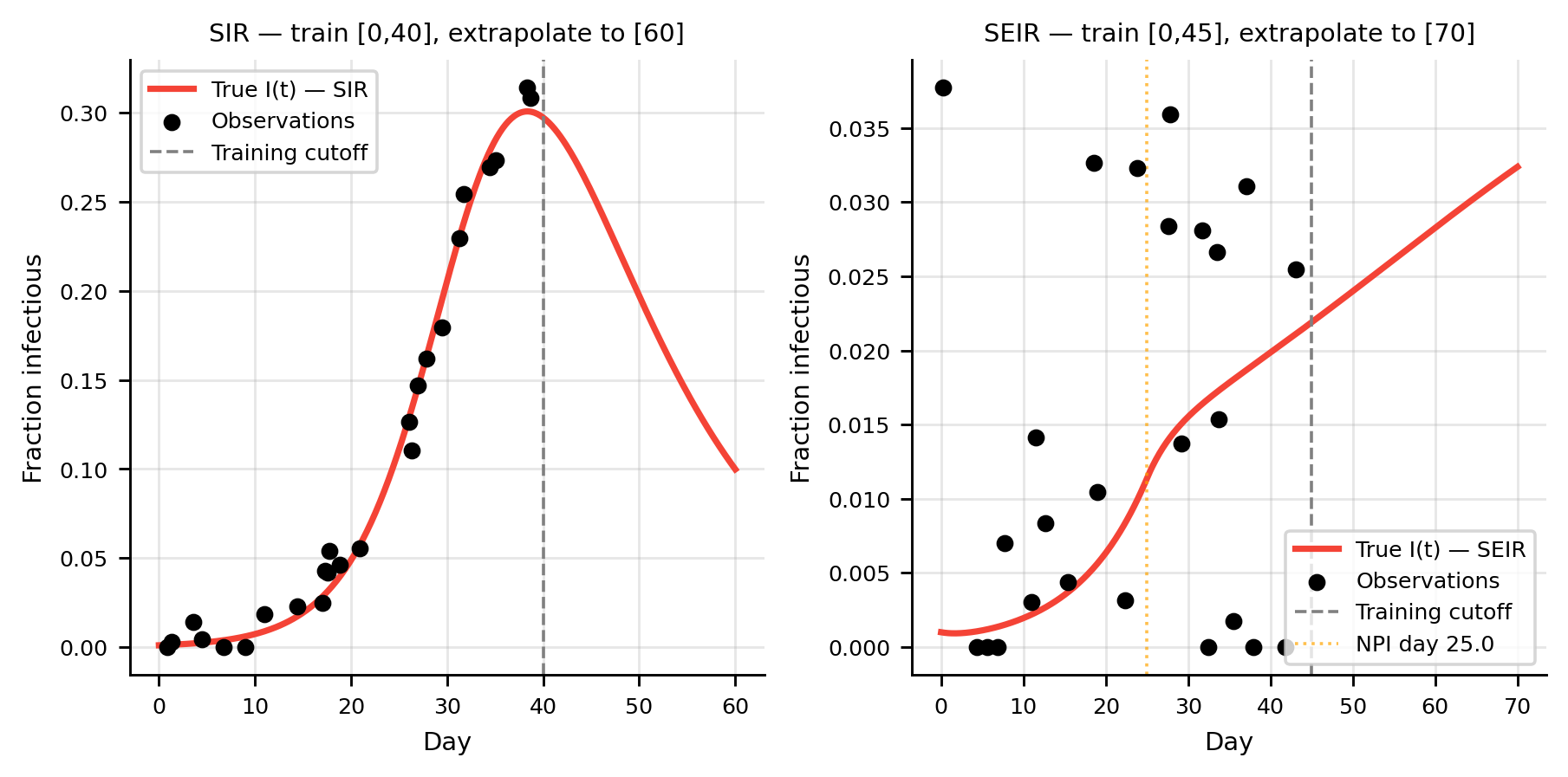
fig, axes = plt.subplots(1, 2, figsize=(7, 3.5))
ax = axes[0]
ax.plot(t_full, sol_sir[:, 1], color=C_I, lw=2, label="True I(t) — SIR")
ax.scatter(t_obs_np, i_noisy, color=C_OBS, s=20, zorder=5, label="Observations")
ax.axvline(T_train, color="gray", lw=1, ls="--", label="Training cutoff")
ax.set(xlabel="Day", ylabel="Fraction infectious",
title=f"SIR — train [0,{int(T_train)}], extrapolate to [{int(T_max)}]")
ax.legend()
ax = axes[1]
ax.plot(t_seir, sol_seir[:, 2], color=C_I, lw=2, label="True I(t) — SEIR")
ax.scatter(t_obs2_np, i_noisy2, color=C_OBS, s=20, zorder=5, label="Observations")
ax.axvline(T_seir_tr, color="gray", lw=1, ls="--", label="Training cutoff")
ax.axvline(NPI_DAY2, color="orange", lw=1, ls=":", alpha=0.7, label=f"NPI day {NPI_DAY2}")
ax.set(xlabel="Day", ylabel="Fraction infectious",
title=f"SEIR — train [0,{int(T_seir_tr)}], extrapolate to [{int(T_seir)}]")
ax.legend()
plt.tight_layout(); plt.show()