SEIR_Euler <- function (params = NULL,
y = NULL,
tbegin = 0,
tend = 1,
dt = 0.2) {
M <- matrix(NA, nrow=(tend-tbegin+1), ncol=length(y)) # output matrix
M[1,] <- y # initial values for the first row
S <- y[1]; E <- y[2]; I <- y[3]; R <- y[4]; CI <- y[5]
N <- S + E + I + R
epsilon <- params[["epsilon"]]
gamma <- params[["gamma"]]
Rt <- params[["Rt"]] # daily reproduction number
for (t in seq(tbegin, tend, by=1)) { # for each day
for (i in seq(dt, 1, dt)) { # sub-intervals that can vary
# beta is already adjusted by N
# t is not an integer
beta <- Rt[floor(t+1+dt)] * gamma # transmission rate
S_to_E <- beta * I * dt
E_to_I <- E * epsilon * dt
I_to_R <- I * gamma * dt
# update state variables
S <- S - S_to_E
E <- E + S_to_E - E_to_I
I <- I + E_to_I - I_to_R
R <- R + I_to_R
CI <- CI + S_to_E
}
# output for each day
M[t+1, 1] <- S
M[t+1, 2] <- E
M[t+1, 3] <- I
M[t+1, 4] <- R
M[t+1, 5] <- CI
}
return(M)
}Estimating the instantaneous reproduction number using the particle filter
A simple particle filter in R
파티클 필터 (particle filter) 를 이용하여 잠재 변수 (latent variable)를 추정하는 과정을 지난 글에서 다루었다. 관찰값들이 코로나 19 일별 감염자일때 감염병 수리 모형을 이용하여 일별 감염재생산지수 (\((R_t\)) 를 추정한다. 아래 글은 Kucharski et al. (1) 논문에 사용되었던 방법을 차용하였다. 파티클 필터 알고리즘의 기초는 Kitagawa (2) 의 방법을 따른다. 이해를 돕기 위해 모형을 단순화 하였고 가상의 데이타를 만들어 내는 과정을 더하였다. 우선 SEIR 모형을 이용해서 가상의 데이타 (일별 감염자 수)를 만든다. 누적 감염자 (cumulative incidence) 를 나타내는 CI라는 변수의 일별 차이를 계산하여 일별 감염자 수를 계산한다. 보통의 SEIR 모형에서는 \(\beta\)가 상수로 취급 되지만 아래 모형에서는 일별 감염 재생산지수 \(R_t = \beta (t) \times D\) \(D\)는 감염 기간)가 방역 정책, 활동 변화 등 이유로 인해 시간에 따라 변화한다고 가정하기 때문에 시간에 따른 함수 \(\beta(t)\)로 표현한다. 우리가 추정 하고자 하는 \(R_t\)를 미리 정의하고 이로 부터 \(\beta(t)\) 를 계산하고 이를 SEIR 모형에 적용하여 가상의 데이타를 만든다.
아래와 같은 방식으로 SEIR 모형을 만든다. 본래 미분식으로 정의하고 deSolve 패키지의 ode 함수 등을 이용하여 적분할 수 있으나 이 글에서는 간단하게 Euler 방법을 사용한다.
일별 감염자 수를 플롯해본다.
# pre-defined Rt
Rt_true <- c(rep(1.2, 15), 0.5*sin(0.1*pi*0:32) + 1.2, rep(0.9, 100))
I0 <- 100 # initially infected people
y0 <- c(S = 1e7-I0, E = 0, I = I0, R = 0, CI = 0) # initial values for state variables
params <- list() # input parameters for the SEIR model
params$Rt <- Rt_true
params$epsilon <- 0.5 # 1/epsilon = latent period
params$gamma <- 0.2 # 1/gamma = duration of infectiousness
tend <- 50 # simulation end time 50 days
res1 <- SEIR_Euler(params = params, y=y0, tend=50) # run the model
res1 <- as.data.frame(res1)
res1$daily_infected <- c(0, diff(res1$V5))
res1$time <- 0:tend
library(ggplot2)
extrafont::loadfonts("win", quiet=TRUE)
theme_set(hrbrthemes::theme_ipsum_rc(base_size=14, subtitle_size=16, axis_title_size=12))
ggplot(res1, aes(x = time, y = daily_infected)) +
geom_line(size = 1.2) +
labs(x = 'Time (day)', y = 'Daily infected')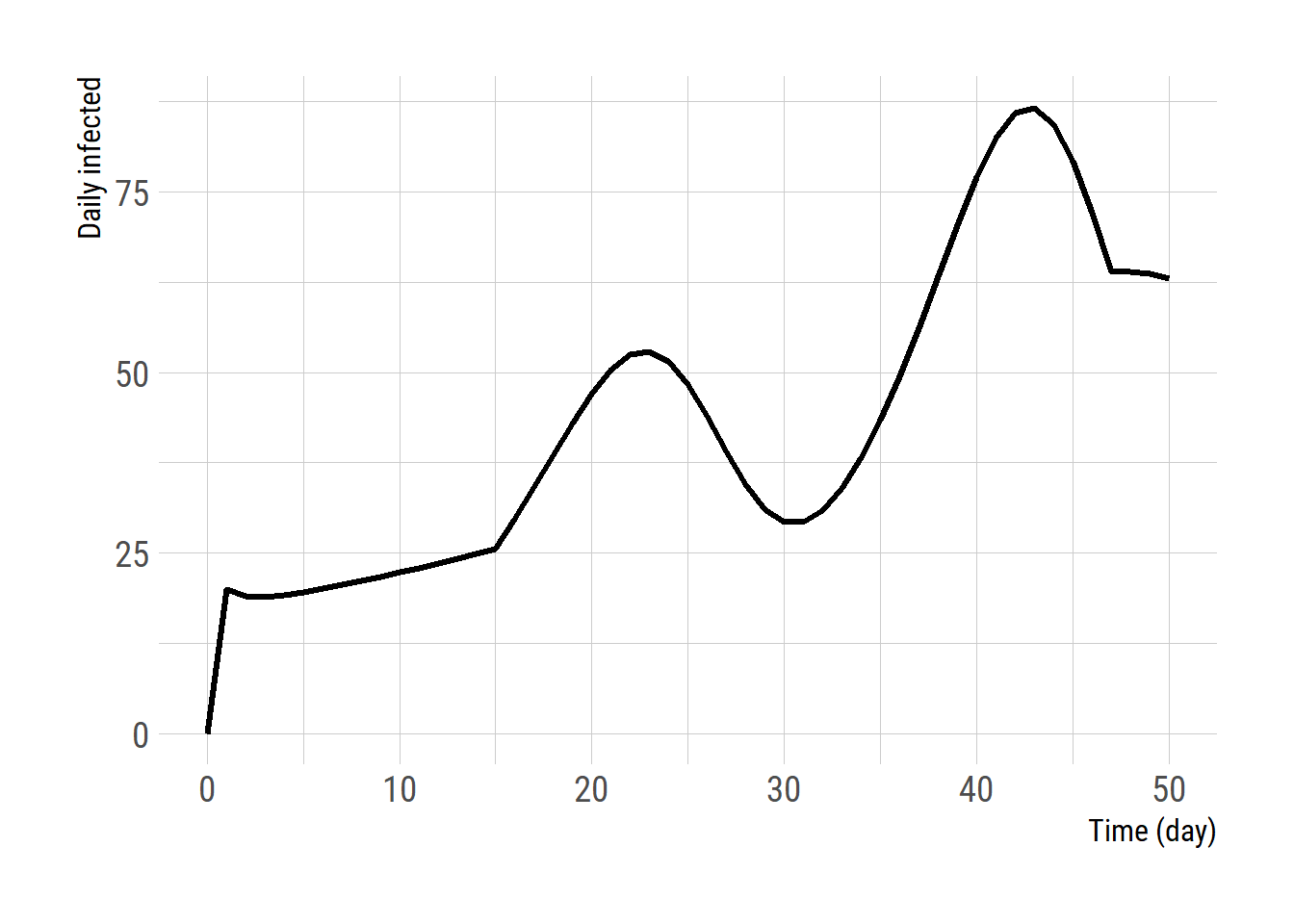
푸아송 분포를 이용하여 가상의 데이타를 만든다.
# Create the data assunming observations are poisson random variable
set.seed(42)
fakedata <- data.frame(daily_infected = rpois(nrow(res1), lambda = res1$daily_infected))일별 변화를 계산하는 SEIR 전파 모형, 행의 수는 파티클 수와 같다.
# stochastic differential equation (with beta(t) moves according to a geometric Brownian motion) are modeled using the Euler-Maruyama method.
# daily change is modeled using the subinterval dt
SEIR_step <- function (params = NULL,
y = NULL,
tbegin = 0,
tend = 1,
dt = 0.2,
beta = NULL) {
# daily infection reset to zero to hold values from tbegin to tend
y[, c("CI")] <- 0
S <- y[, "S"]
E <- y[, "E"]
I <- y[, "I"]
R <- y[, "R"]
daily_infected <- y[, "CI"]
N <- S + E + I + R
epsilon <- params[["epsilon"]]
gamma <- params[["gamma"]]
for (i in seq((tbegin + dt), tend, dt)) {
# beta is already assumed to be adjusted by N such that it can
# be translated to Rt by multiplying the duration of infectiousness
S_to_E <- beta * I * dt
E_to_I <- E * epsilon * dt
I_to_R <- I * gamma * dt
# Process model for SEIR
S <- S - S_to_E
E <- E + S_to_E - E_to_I
I <- I + E_to_I - I_to_R
R <- R + I_to_R
daily_infected <- daily_infected + S_to_E
}
y[, "S"] <- S
y[, "E"] <- E
y[, "I"] <- I
y[, "R"] <- R
y[, "CI"] <- daily_infected
return(y)
}파티클 필터링 함수
pfilter <- function (params, # parameters
y, # initial values of state variables
data, # input data set
npart = 1000, # number of particles
tend = NULL, # simulation stop time
dt = 0.2) {
# Assumptions - using daily growth rate
nstatevar <- length(y) # number of state variables
if(is.null(tend)) {
tend = nrow(data)
}
# to store state variables
latent_var <- array(0,
dim = c(npart, tend, nstatevar),
dimnames = list(NULL, NULL, names(y)))
# latent_var[, 1, ] <- y
for (nm in names(y)) { # initial value
latent_var[, 1, nm] <- y[[nm]]
}
## parameters
gamma <- params[["gamma"]]
beta0 <- params[["R0"]] * gamma
beta_sd <- params[["betavol"]]
beta <- matrix(rnorm(npart * tend, mean = 0, sd = beta_sd), nrow = tend)
beta[1,] <- beta0 # this is updated at t=2
wt <- matrix(NA, nrow = npart, ncol = tend) # weight (likelihood)
wt[, 1] <- 1 / npart # initial weights
W <- matrix(NA, nrow = npart, ncol = tend) # normalized weights
A <- matrix(NA, nrow = npart, ncol = tend) # Resample according to the normalized weight
for (t in 2:tend) {# begin particle loop
# beta changes according to a Geometric Brownian motion
beta[t, ] <- beta[t-1, ] * exp(beta[t, ])
# run process model
latent_var[, t, ] <- SEIR_step(params = params,
y = latent_var[, t-1, ],
tbegin = t-1,
tend = t,
dt = dt,
beta = beta[t,])
# calculate weights (likelihood)
# wt[, t] <- assign_weights(var = latent_var, t = t, data = data)
case_expected <- latent_var[, t, "CI"]
case_data <- round(unlist(data[t, "daily_infected"]))
expected_val <- pmax(0, case_expected) # make sure that the value is not negative
log_lik <- dpois(round(case_data), lambda = expected_val, log = T)
wt[, t] <- exp(log_lik)
# normalize particle weights
W[, t] <- wt[, t] / sum(wt[, t])
# resample particles by sampling parent particles according to weights
A[, t] <- sample(1:npart, prob = W[1:npart, t], replace = T)
# Resample particles for corresponding variables
latent_var[, t,] <- latent_var[A[, t], t,]
beta[t,] <- beta[t, A[, t]] #- needed for random walk on beta
} # end particle loop
# Marginal likelihoods
lik_values <- rep(NA, tend)
for (t in 1:tend) {
lik_values[t] <- log(sum(wt[1:npart, t])) # log-likelihoods
}
# averaged log likelihoods log(L/(npart^tend))
loglik <- - tend * log(npart) + sum(lik_values)
return (list(lik_marginal = lik_values,
lik_overall_average = loglik,
latent_var_filtered = latent_var,
beta_filtered = beta,
W = W, A = A))
}일별 변화를 계산하는 SEIR 전파 모형, 행의 수는 파티클 수와 같다.
params$R0 <- 2
params$betavol <- 0.3
sample <- pfilter(params=params, # parameters
y=y0, # initial values of state variables
data=fakedata, # input data set
npart = 1000, # number of particles
tend = tend, # simulation stop time
dt = 0.2)
observed <- fakedata$daily_infected[2:nrow(fakedata)]Plot the results
# draw incidence plot
daily_inc_summary <- t(apply(sample$latent_var_filtered[,,5], 2, quantile,
probs=c(0.025, 0.5, 0.975)))
df <- cbind(data.frame(time=1:(nrow(res1)-1), observed = observed), daily_inc_summary)
ggplot(df, aes(x=time)) +
geom_ribbon(aes(ymin=`2.5%`, ymax=`97.5%`), fill="steelblue", alpha=0.8)+
geom_line(aes(y=`50%`), color="steelblue")+
geom_point(aes(y=observed), color = "darkred")+
labs(x="Time", y="Daily incidence")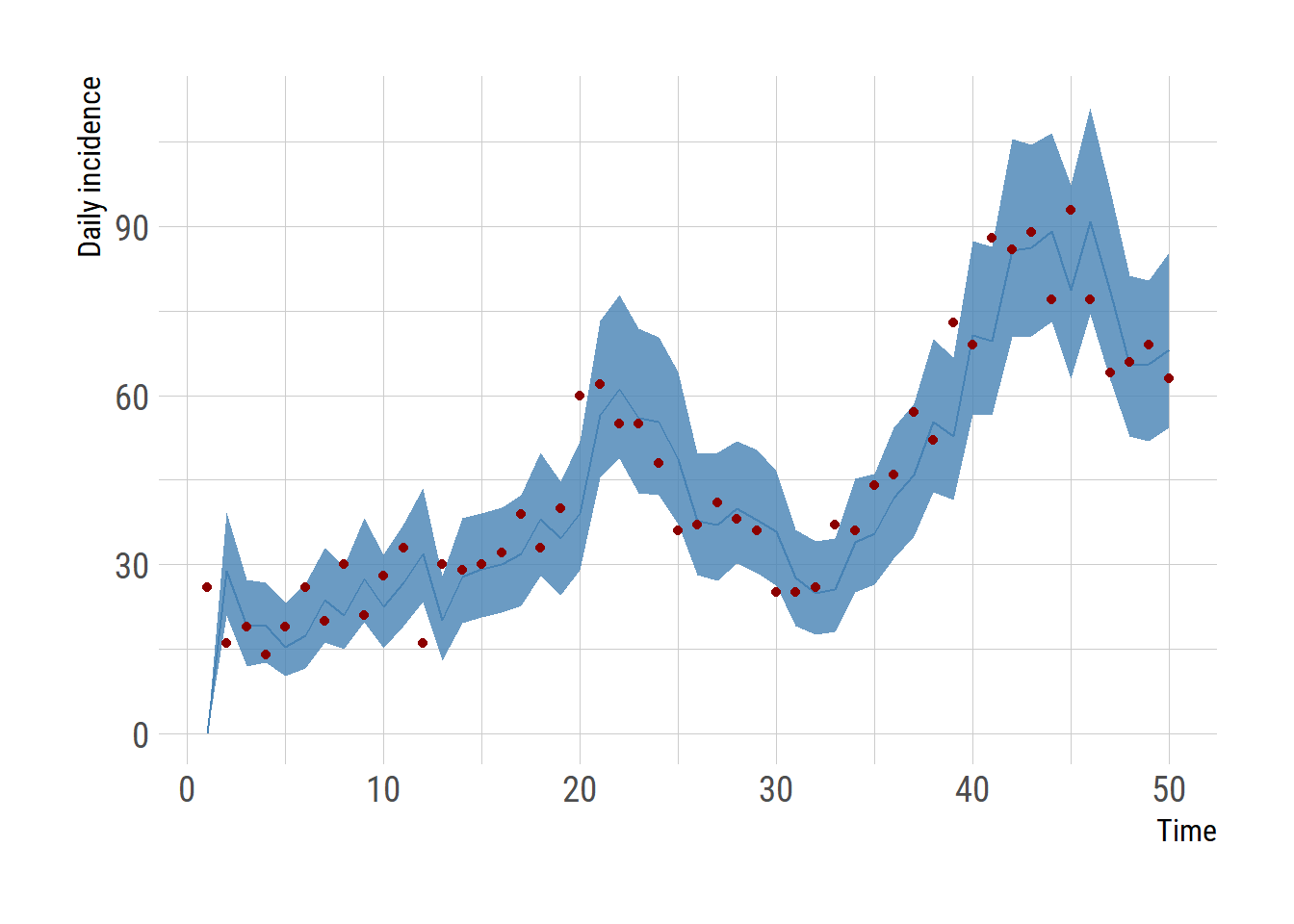
# draw daily Rt plot
dur <- 1/params$gamma
daily_Rt_summary <- t(apply(sample$beta_filtered * dur, 1, quantile,
probs=c(0.025, 0.5, 0.975)))
df <- cbind(data.frame(time=1:(nrow(res1)-1), true_Rt = Rt_true[2:51]), daily_Rt_summary)
ggplot(df, aes(x=time)) +
geom_ribbon(aes(ymin=`2.5%`, ymax=`97.5%`), fill="darkgreen", alpha=0.7)+
geom_line(aes(y=`50%`), color="darkgreen")+
geom_point(aes(y=true_Rt), color = "black") +
labs(x="Time", y=expression(italic(R[t])))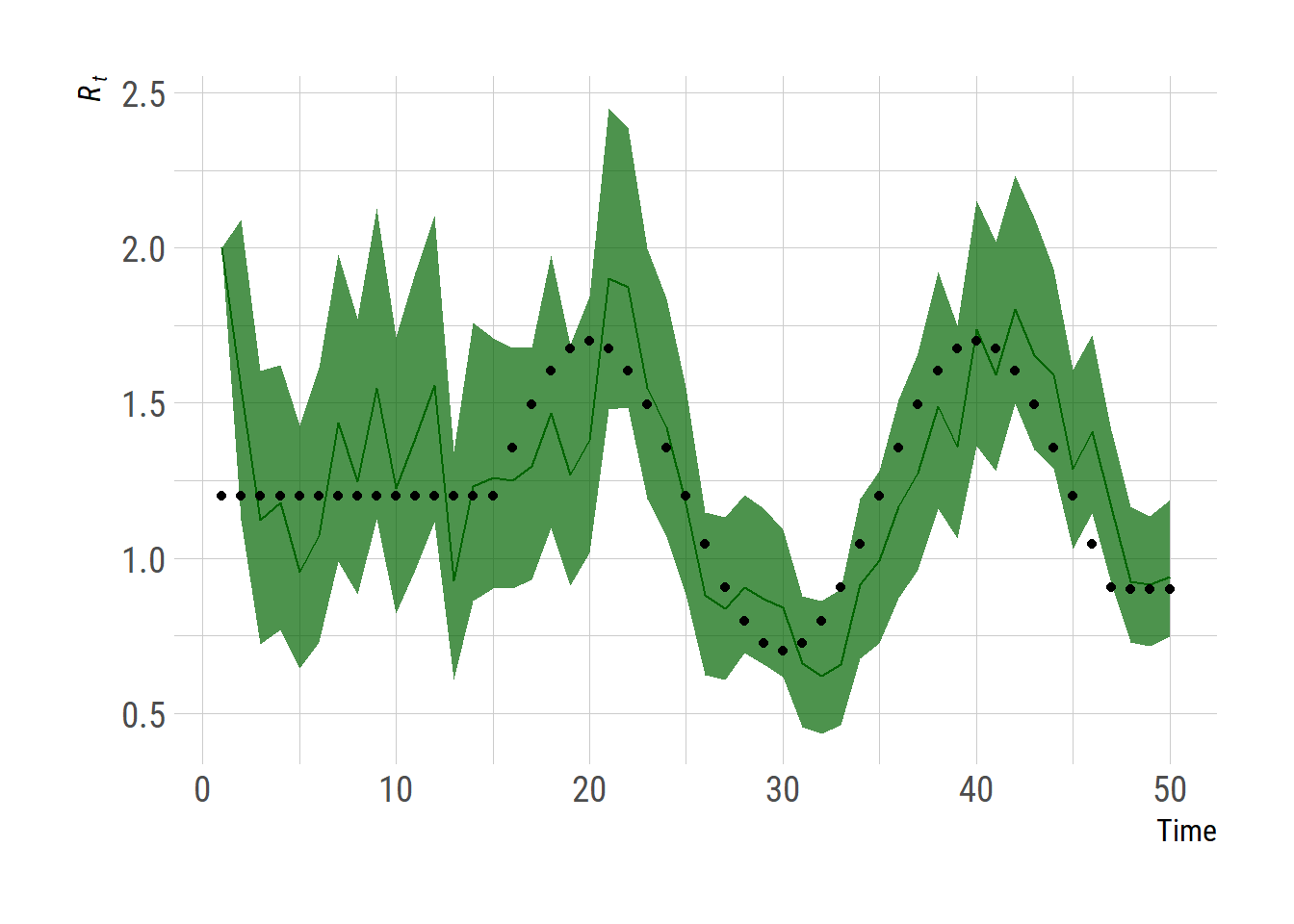
# ggsave("particle_filter_covid.png", gg, units="in", width=3.4*2, height=2.7*2)