---
title: "Digital Twins in Clinical Trials"
subtitle: "From virtual patients to immune simulators — how computational replicas are reshaping vaccine development"
author: "Jong-Hoon Kim"
date: "2026-04-21"
categories: [digital twin, clinical trial, vaccine, infectious disease, R]
bibliography: references.bib
csl: https://www.zotero.org/styles/vancouver
format:
html:
toc: true
toc-depth: 3
code-fold: true
code-tools: true
number-sections: true
fig-width: 7
fig-height: 4.5
fig-align: center
knitr:
opts_chunk:
message: false
warning: false
fig.retina: 2
---
```{r setup, include=FALSE}
library(deSolve)
library(ggplot2)
library(dplyr)
library(tidyr)
library(patchwork)
```
# What is a digital twin?
The term *digital twin* was coined by Michael Grieves around 2002 in a manufacturing context: a real artefact, a virtual replica that mirrors it, and a bidirectional data connection keeping the two in sync [@grieves2017]. In aerospace and civil engineering the idea is now mature — you build a digital twin of a turbine blade or a bridge, run it under load in simulation, and intervene on the physical object only when the model flags a problem.
Medicine arrived at the concept via a different route. As electronic health records, multi-omics assays, and wearable sensors began generating continuous streams of data about individual patients, it became conceivable to build a *patient-level* model rather than a population-average one: a computational instance calibrated to one person's physiology that could be queried, perturbed, and iterated faster than any wet-lab experiment. Clinical decision support, drug dosing, and oncology treatment planning were early targets [@bjornsson2020; @stahlberg2022].
Clinical trials entered the picture when researchers realised that a well-validated patient model could play the role of a *counterfactual control*: what would have happened to this patient had they received placebo? If that counterfactual is reliable enough, it can supplement — or in limited circumstances replace — a randomised control arm. The potential benefits are compelling:
- **Smaller trials**: fewer participants needed when the control response is already known from simulation.
- **Greater equity**: patients who would otherwise receive only placebo may instead receive active treatment.
- **Personalised outcomes**: population-average efficacy estimates give way to individual trajectory predictions.
- **Faster iteration**: many candidate regimens are eliminated in silico before any needle is lifted.
This post reviews how digital twins are being built for infectious-disease vaccine trials, what the underlying modelling machinery looks like, and where the major unresolved challenges lie.
---
# The modelling stack
A clinical-trial digital twin is rarely one model. It is a *stack* of coupled models operating at different biological scales [@masison2021]:
| Scale | Examples | Key state variables |
|---|---|---|
| Molecular | Receptor binding, antigen processing | Antibody titres, antigen load |
| Cellular | T-cell, B-cell, innate cell dynamics | Cell counts, activation states |
| Tissue / organ | Lymph node germinal centre, lung epithelium | Local cytokine concentrations |
| Individual | Systemic immune kinetics, PK/PD | Viral load, IgG, temperature |
| Population | Epidemic transmission | S, E, I, R compartments |
For vaccine trials, the most consequential scales are the *individual* and the *population*. Individualisation distinguishes a digital twin from a conventional pharmacometric model; population coupling connects immune responses to clinical endpoints (does this person get infected? are they hospitalised?).
---
# Immune digital twins
## Quantitative systems pharmacology
Quantitative systems pharmacology (QSP) sits at the individual scale and is currently the dominant modelling framework for immune digital twins in pharma [@desikan2024]. A QSP model is a large ODE system encoding, mechanistically, how a vaccine depot is processed by dendritic cells, how antigen-specific B cells expand in germinal centres, how antibody titres rise and wane, and how cross-reactivity bridges strain variants.
For mRNA vaccines, the model must also capture mRNA clearance, spike protein expression kinetics, and toll-like receptor-mediated innate activation at the injection site. Dasti et al. [@dasti2025] built a multiscale QSP model for Pfizer and Moderna vaccines, calibrating dendritic-cell recruitment, monocyte activation, and antibody kinetics against clinical immunogenicity data. The result is a *virtual patient cohort*: a distribution of parameter sets, each plausible biologically, that spans the observed inter-individual variability in antibody response.
## The UISS platform for COVID-19 vaccines
One of the first published in-silico vaccine trials for a novel pathogen used the Universal Immune System Simulator (UISS), an agent-based model that tracks individual immune cells, antigens, and cytokines on a spatial grid [@kutovyi2020]. The COVID-19 application simulated three candidate vaccine strategies — different antigen doses and prime-boost schedules — and ranked them by predicted neutralising-antibody titre. The simulations were completed before any Phase 1 data were available, providing hypothesis-ranked predictions that were later tested experimentally.
UISS represents one extreme of the modelling spectrum: computationally expensive, mechanistically granular, but hard to calibrate to individual patients. QSP models occupy the other extreme: ODE-based, fast to run, but aggregating cellular processes into lumped rate constants. Hybrid approaches that embed reduced-order immune modules inside epidemic transmission models are an active research frontier [@hartmann2024].
---
# Synthetic control arms
## The idea
A *synthetic control arm* (SCA) is a statistical construction: a set of simulated or historically matched patients who stand in for the placebo group in a single-arm trial. The digital-twin framing is strongest when the counterfactual is generated by a calibrated mechanistic model rather than a propensity-score-matched historical cohort, but both use cases appear in the regulatory literature.
The appeal is clearest in settings where a placebo arm is ethically difficult (pandemic vaccines with high attack rates), practically impossible (rare diseases, paediatric populations), or commercially unattractive (sponsors prefer all active-arm data). Pammi et al. [@landig2025] review how digital twins could address precisely these challenges in paediatric infectious-disease trials.
## Regulatory position
The FDA's Model-Informed Drug Development (MIDD) program actively encourages submission of modelling evidence alongside clinical data, and has issued paired-meeting guidance for sponsors wanting to use quantitative models in regulatory submissions [@fda2020midd]. The EMA qualified PROCOVA — a prognostic-covariate adjustment method that uses a digital-twin-derived prognostic score to reduce variance in primary endpoints — giving European sponsors a concrete, accepted implementation path [@janssen2025]. ICH M15, released in draft in November 2024, provides the first international harmonised guidance on model-informed drug development.
Synthetic control arms do not yet replace randomised placebo groups in pivotal trials for most indications. The evidentiary bar is a *validated predictive model*: the sponsor must demonstrate, typically using held-out historical data, that the model's counterfactual trajectories match what actually happened to control patients. The challenge is especially sharp for novel pathogens where no historical controls exist.
---
# Vaccine trials for infectious diseases
## COVID-19
The COVID-19 pandemic stress-tested the entire paradigm. Vaccine trials of unprecedented speed (under a year from sequence to efficacy readout) relied heavily on existing correlates-of-protection models. Bordukova et al. [@bhatnagar2022] describe how generative AI accelerated the construction of in-silico trial arms, compressing what would have been multi-year simulation programmes into months.
Independently, mathematical models of SARS-CoV-2 transmission and spread [@meehan2020] were coupled to trial simulation frameworks to predict vaccine efficacy across age strata and variant scenarios. The key question these models addressed — *what neutralising-antibody titre predicts protection?* — remains central to all COVID-19 booster policy.
## Influenza
Influenza presents a different challenge: annual antigenic drift means the vaccine must be updated each season, and efficacy varies enormously depending on how well the vaccine strain matches the circulating strain. Digital twins for influenza trials need to represent *strain-specific* immune memory — accounting for the imprinting effect of childhood infections, cross-reactive responses, and waning from prior vaccination [@hartmann2024].
A physics-informed neural network approach [@raissi2019] could, in principle, embed a reduced SEIR-type transmission model inside an individual-patient simulator, jointly calibrating epidemic parameters and individual immune parameters from longitudinal cohort data. This kind of *universal differential equation* framework [@rackauckas2020] is gaining traction as the methodological bridge between mechanistic models and machine-learning flexibility.
## Wearable sensors as real-time digital-twin inputs
Steinhubl et al. [@grieff2025] demonstrated a particularly direct implementation of the digital-twin concept for vaccine trials: a wearable torso patch records continuous physiological signals (heart rate variability, skin temperature, respiratory rate) in 88 participants receiving 104 vaccine doses across different products. A machine-learning similarity model constructs each participant's *individualised physiological baseline* — effectively a personalised digital twin of their resting physiology — and expresses post-vaccination deviation against that twin. The resulting personalised inflammatory biomarker correlated significantly with serum CRP and IFN-γ, markers of vaccine-induced reactogenicity that normally require blood draws. This is real-time digital-twin feedback at human scale.
---
# A minimal illustration: SIR with vaccine efficacy
To ground the ideas concretely, consider the simplest possible coupling between an individual immune response and a population-level trial endpoint. Suppose the vaccine produces a neutralising-antibody titre $T$ in individual $i$, drawn from a log-normal distribution calibrated from immunogenicity data. Protection from infection is a sigmoidal function of titre:
$$
\text{VE}_i = \frac{T_i^k}{T_i^k + \text{EC}_{50}^k}
$$
where $\text{EC}_{50}$ is the titre at 50% protection and $k$ is a Hill coefficient. We can propagate this through a two-arm SIR model (vaccinated arm with heterogeneous VE, placebo arm) to simulate an attack-rate trial.
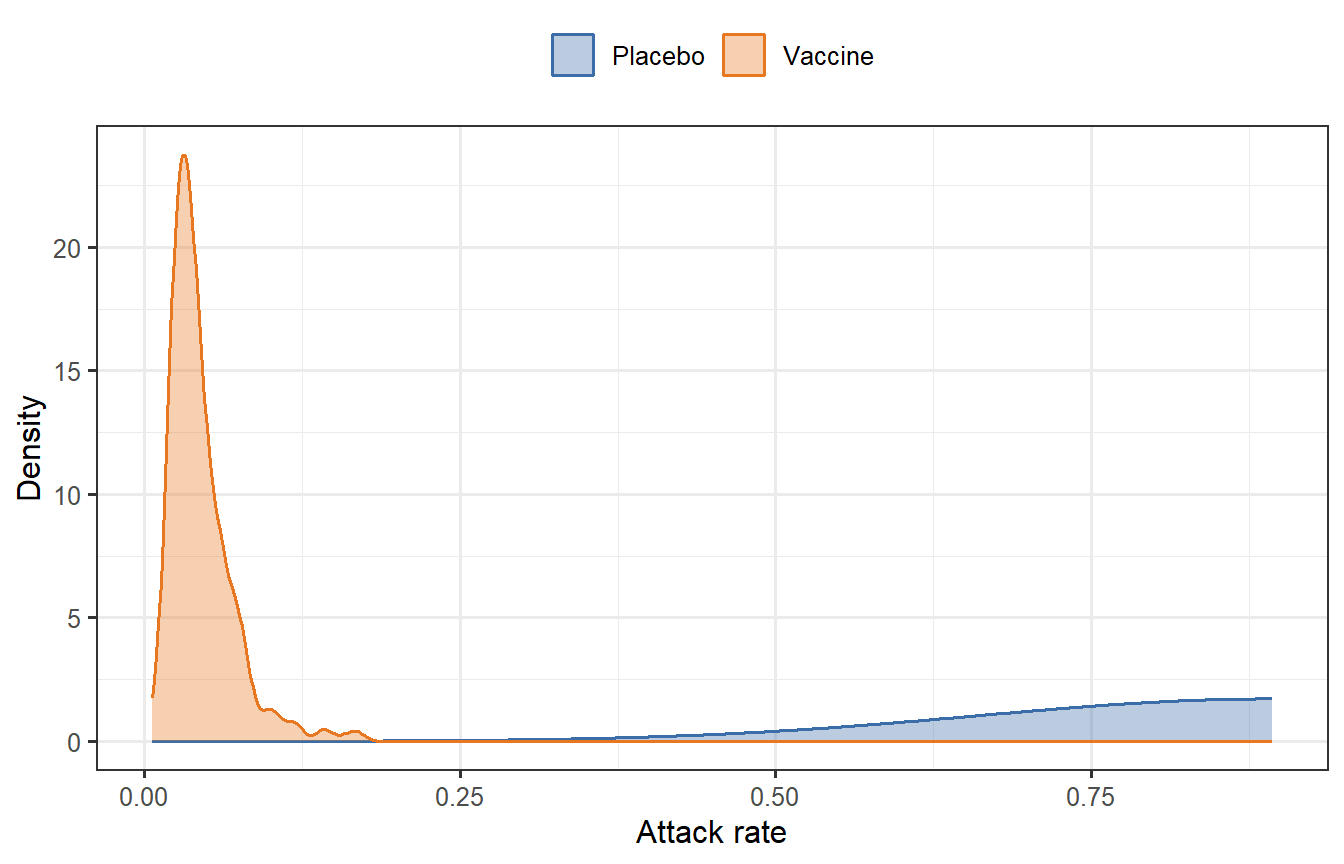
```{r sir-dt, fig.cap="Simulated vaccine trial: attack-rate distributions under heterogeneous individual-level VE (Hill model). The vaccine arm (orange) shows substantially fewer infections than placebo (blue) but with considerable individual variability — the sort of prediction a digital twin generates before a trial is run."}
set.seed(42)
n <- 2000
# Titre distribution (log-normal)
mu_log <- log(100)
sigma_log <- 0.8
Ti <- rlnorm(n, meanlog = mu_log, sdlog = sigma_log)
# Hill model for VE
EC50 <- 80; k <- 2
VEi <- Ti^k / (Ti^k + EC50^k)
# SIR for each arm
sir_attack <- function(R0, VE_vec, N = 1000, I0 = 1, dt = 0.1, tmax = 200) {
n_ind <- length(VE_vec)
# Mean VE for the arm
mean_beta_reduction <- mean(1 - VE_vec)
S <- N - I0; I <- I0; R <- 0
gamma <- 0.1
beta0 <- R0 * gamma / N
beta <- beta0 * mean_beta_reduction
t <- 0
while (I > 0.5 && t < tmax) {
dS <- -beta * S * I
dI <- beta * S * I - gamma * I
dR <- gamma * I
S <- S + dS * dt
I <- I + dI * dt
R <- R + dR * dt
t <- t + dt
}
R / N # attack rate
}
# Bootstrap trial simulations
n_trials <- 500
R0 <- 2.5
# Vaccinated arm: random subset of virtual patients
vacc_ar <- replicate(n_trials, {
idx <- sample(n, 200, replace = FALSE)
sir_attack(R0, VEi[idx])
})
# Placebo arm: VE = 0
placebo_ar <- replicate(n_trials, {
sir_attack(R0, rep(0, 200))
})
df_plot <- bind_rows(
tibble(arm = "Vaccine", attack_rate = vacc_ar),
tibble(arm = "Placebo", attack_rate = placebo_ar)
)
ggplot(df_plot, aes(x = attack_rate, fill = arm, colour = arm)) +
geom_density(alpha = 0.35, linewidth = 0.7) +
scale_fill_manual(values = c("Vaccine" = "#E87722", "Placebo" = "#3B6EA8")) +
scale_colour_manual(values = c("Vaccine" = "#E87722", "Placebo" = "#3B6EA8")) +
labs(
x = "Attack rate",
y = "Density",
fill = NULL, colour = NULL
) +
theme_bw(base_size = 12) +
theme(legend.position = "top")
```
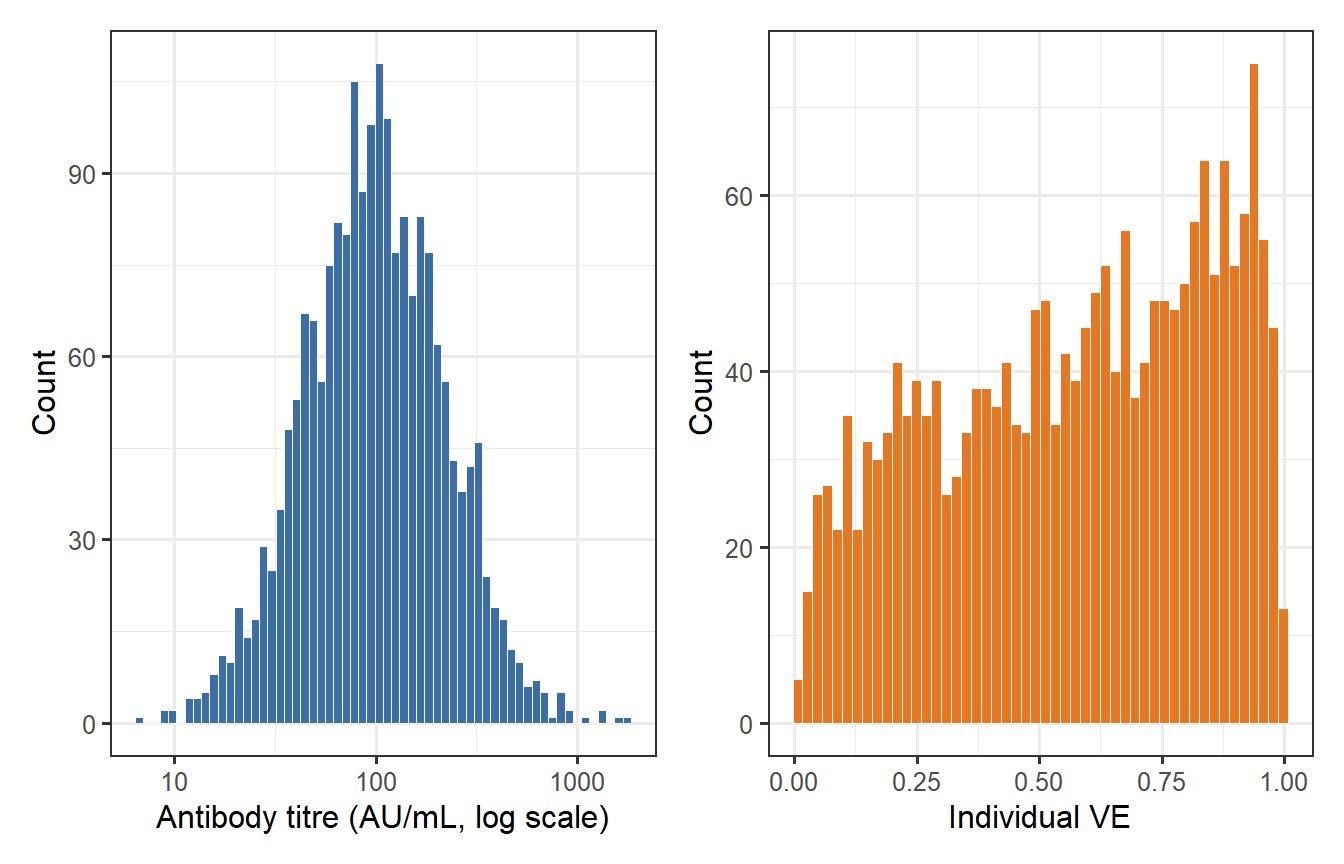
```{r ve-distribution, fig.cap="Distribution of individual vaccine efficacy (VE) arising from log-normal antibody titre variability and a Hill protection model (EC50 = 80, k = 2). Even a vaccine with mean VE ~ 0.85 leaves a long left tail of poorly protected individuals — exactly the heterogeneity a digital twin must capture to inform trial design."}
df_ve <- tibble(titre = Ti, VE = VEi)
p1 <- ggplot(df_ve, aes(x = titre)) +
geom_histogram(bins = 60, fill = "#3B6EA8", colour = "white", linewidth = 0.2) +
scale_x_log10() +
labs(x = "Antibody titre (AU/mL, log scale)", y = "Count") +
theme_bw(base_size = 12)
p2 <- ggplot(df_ve, aes(x = VE)) +
geom_histogram(bins = 50, fill = "#E87722", colour = "white", linewidth = 0.2) +
labs(x = "Individual VE", y = "Count") +
theme_bw(base_size = 12)
p1 + p2
```
The simulation illustrates three features that make digital twins useful for trial design:
1. **Heterogeneity**: even a good vaccine leaves a tail of poorly protected individuals. Conventional trial design based on mean VE obscures this.
2. **Counterfactual control**: once the vaccine arm's VE distribution is estimated from immunogenicity data, the placebo attack-rate distribution can be simulated without actually randomising patients to placebo.
3. **Sensitivity analysis**: changing $\text{EC}_{50}$, $k$, or the titre distribution immediately produces a new attack-rate prediction — useful for go/no-go decisions early in development.
---
# Challenges and limitations
## Model validation
A digital twin's predictions are only as trustworthy as its calibration data. For established vaccines with decades of immunogenicity and efficacy data (inactivated influenza, yellow fever 17D), calibration is tractable. For novel antigens against emerging pathogens, the twin must be built and used before most of the data it needs exist — a fundamental bootstrap problem. Prospective validation on held-out cohorts and pre-specified decision criteria are essential [@kleeberger2025].
## Individual vs population parameters
QSP models routinely have hundreds of parameters. The number of identifiable parameter combinations from typical Phase 1 immunogenicity data (n = 30–60, a handful of antibody timepoints) is far smaller. Virtual patient cohorts are often generated by Latin hypercube sampling of plausible parameter ranges [@hartmann2024], but the realism of the resulting distribution depends entirely on the quality of prior biological knowledge used to define those ranges.
## Regulatory acceptance
Despite FDA and EMA progress, no pivotal vaccine trial has yet received primary approval on the basis of a synthetic control arm alone. The regulatory framework treats modelling evidence as supportive — capable of reducing sample size or justifying extrapolation — rather than as a primary evidentiary source. Akbarialiabad et al. [@janssen2025] lay out a roadmap for trials that explicitly pre-register their digital-twin components alongside conventional statistical analysis plans.
## Equity and bias
Virtual patient cohorts built from historically skewed clinical databases may underrepresent populations that were already underrepresented in the underlying data — amplifying, rather than correcting, health equity gaps. Pammi et al. [@landig2025] identify this as an especially acute concern for paediatric trials, where the data scarcity that motivates digital twins also limits their calibration.
## Immune waning and long-horizon prediction
Most immune digital twins are validated against short-term endpoints (peak antibody titre at 28 days). Long-horizon predictions — what is this person's protection 18 months after a third dose? — involve waning kinetics that are poorly characterised and likely depend on mechanisms (memory B-cell maintenance, T-follicular helper cell longevity) not represented in current ODE-based models.
---
# Outlook
The convergence of three trends suggests digital twins will become structural components of vaccine development over the next decade:
1. **Richer individual-level data.** Continuous wearable monitoring [@grieff2025], single-cell immune profiling, and multi-omic biobanks are steadily providing the calibration data that digital twins need. The gap between what models can represent and what data can constrain is narrowing.
2. **Physics-informed machine learning.** Methods that embed known biology (ODE constraints, mass-balance equations) into neural architectures [@raissi2019; @rackauckas2020] allow flexible individual-level fitting without sacrificing mechanistic interpretability. This is the direction the field is moving for complex multi-scale problems.
3. **Regulatory confidence-building.** The ICH M15 harmonisation, EMA PROCOVA qualification, and FDA MIDD program are creating precedents that sponsors can cite. As successful examples accumulate, the bar for using digital-twin evidence in regulatory submissions will lower.
For infectious-disease vaccine trials specifically, the most tractable near-term application is **adaptive trial design**: a digital twin informs interim decisions (dose, schedule, subgroup enrichment) rather than replacing the trial's placebo arm. This requires only that the model is predictive within the trial — a much weaker requirement than the full counterfactual use case — and it preserves randomisation, which regulators still regard as the gold standard for causal inference.
The more ambitious goal — a fully *in silico* Phase 2 proof-of-concept for a novel antigen, with a synthetic control arm accepted by regulators as the primary comparison — remains 5–10 years away for most vaccine platforms. It will require pathogen-specific immune models of demonstrated external validity, open data-sharing infrastructure, and a generation of regulators and trial statisticians trained in model assessment. Each of those prerequisites is in active development.
---
# References {.unnumbered}
::: {#refs}
:::
```{r session-info, echo=FALSE}
sessionInfo()
```