# SEIR model returning peak infectious count
seir_peak <- function(beta, sigma, gamma,
N = 100000, I0 = 50, days = 120) {
S <- N - I0; E <- 0; I <- I0; R <- 0
peak <- I
for (d in seq_len(days)) {
new_e <- beta * S * I / N
new_i <- sigma * E
new_r <- gamma * I
S <- S - new_e; E <- E + new_e - new_i
I <- I + new_i - new_r; R <- R + new_r
if (I > peak) peak <- I
}
peak / N # return as fraction
}Global Sensitivity Analysis as a Billable Model Audit Service
Which parameters drive your epidemic forecast? Sobol indices and Morris screening turn model uncertainty into a professional deliverable. Skill 2 of 20.
1 The business case
A health department client asks: “You say \(R_0 = 1.8\), but how sensitive is your 14-day forecast to that estimate? What if it’s actually 2.2?” Without sensitivity analysis you can only shrug. With it, you can hand them a one-page report quantifying exactly how much each uncertain parameter moves the forecast needle.
Global sensitivity analysis (SA) (1,2) answers: which inputs explain most of the output variance? This is a different question from “what happens if I change \(\beta\) by 10%” (local sensitivity). Global SA samples the entire input space and decomposes variance into contributions from each parameter and their interactions.
It is also a distinct, defensible deliverable — a model audit that commands consulting fees separate from the core subscription.
2 Variance-based (Sobol) indices
For a model \(Y = f(X_1, \ldots, X_k)\), the first-order Sobol index for parameter \(X_i\) is:
\[S_i = \frac{V_{X_i}[\,E_{\mathbf{X}_{\sim i}}(Y \mid X_i)\,]}{V(Y)}\]
It measures the fraction of output variance explained by \(X_i\) alone. The total-order index \(S_{T_i}\) captures \(X_i\)’s contribution including all interactions with other parameters:
\[S_{T_i} = 1 - \frac{V_{X_{\sim i}}[\,E_{X_i}(Y \mid \mathbf{X}_{\sim i})\,]}{V(Y)}\]
When \(S_{T_i} \gg S_i\), interactions dominate. Both indices lie in \([0,1]\) and \(\sum_i S_i \leq 1\) (3).
3 Implementing Sobol indices in R
We compute Sobol indices for peak infected count in an SEIR model with three uncertain inputs: \(\beta\), \(\sigma\) (incubation rate), and \(\gamma\) (recovery rate).
set.seed(2024)
# Parameter ranges (uniform priors)
ranges <- list(
beta = c(0.20, 0.55),
sigma = c(1/8, 1/3),
gamma = c(1/10, 1/5)
)
# Saltelli (2010) estimator needs two independent base samples A, B
n <- 4000
A <- sapply(ranges, function(r) runif(n, r[1], r[2]))
B <- sapply(ranges, function(r) runif(n, r[1], r[2]))
# Evaluate model on A and B
yA <- mapply(seir_peak, A[,"beta"], A[,"sigma"], A[,"gamma"])
yB <- mapply(seir_peak, B[,"beta"], B[,"sigma"], B[,"gamma"])
# For each parameter i, create AB_i: A with column i replaced by B
sobol_indices <- function(yA, yB, A, B, ranges) {
k <- ncol(A)
nms <- colnames(A)
S <- numeric(k); ST <- numeric(k)
Vy <- var(c(yA, yB))
for (i in seq_len(k)) {
AB_i <- A
AB_i[, i] <- B[, i]
yABi <- mapply(seir_peak,
AB_i[,"beta"], AB_i[,"sigma"], AB_i[,"gamma"])
# Jansen estimator (more robust than Saltelli original)
S[i] <- (var(yB) - mean((yB - yABi)^2) / 2) / Vy
ST[i] <- mean((yA - yABi)^2) / (2 * Vy)
}
data.frame(parameter = nms,
S1 = pmax(0, S),
ST = pmax(0, ST))
}
idx <- sobol_indices(yA, yB, A, B, ranges)
print(idx) parameter S1 ST
1 beta 0.55765633 0.56522359
2 sigma 0.04961737 0.04964628
3 gamma 0.40615158 0.41383019library(ggplot2)
idx_long <- rbind(
data.frame(parameter = idx$parameter, index = "First-order (S1)",
value = idx$S1),
data.frame(parameter = idx$parameter, index = "Total-order (ST)",
value = idx$ST)
)
ggplot(idx_long, aes(x = reorder(parameter, -value),
y = value, fill = index)) +
geom_col(position = "dodge", width = 0.6) +
scale_fill_manual(values = c("First-order (S1)" = "steelblue",
"Total-order (ST)" = "firebrick"),
name = NULL) +
geom_hline(yintercept = 0.05, linetype = "dashed", colour = "grey40") +
annotate("text", x = 0.6, y = 0.07,
label = "5% threshold", hjust = 0, size = 3.2, colour = "grey40") +
labs(x = "Parameter", y = "Sobol index",
title = "Global sensitivity: SEIR peak infectious fraction") +
theme_minimal(base_size = 13) +
theme(legend.position = "top")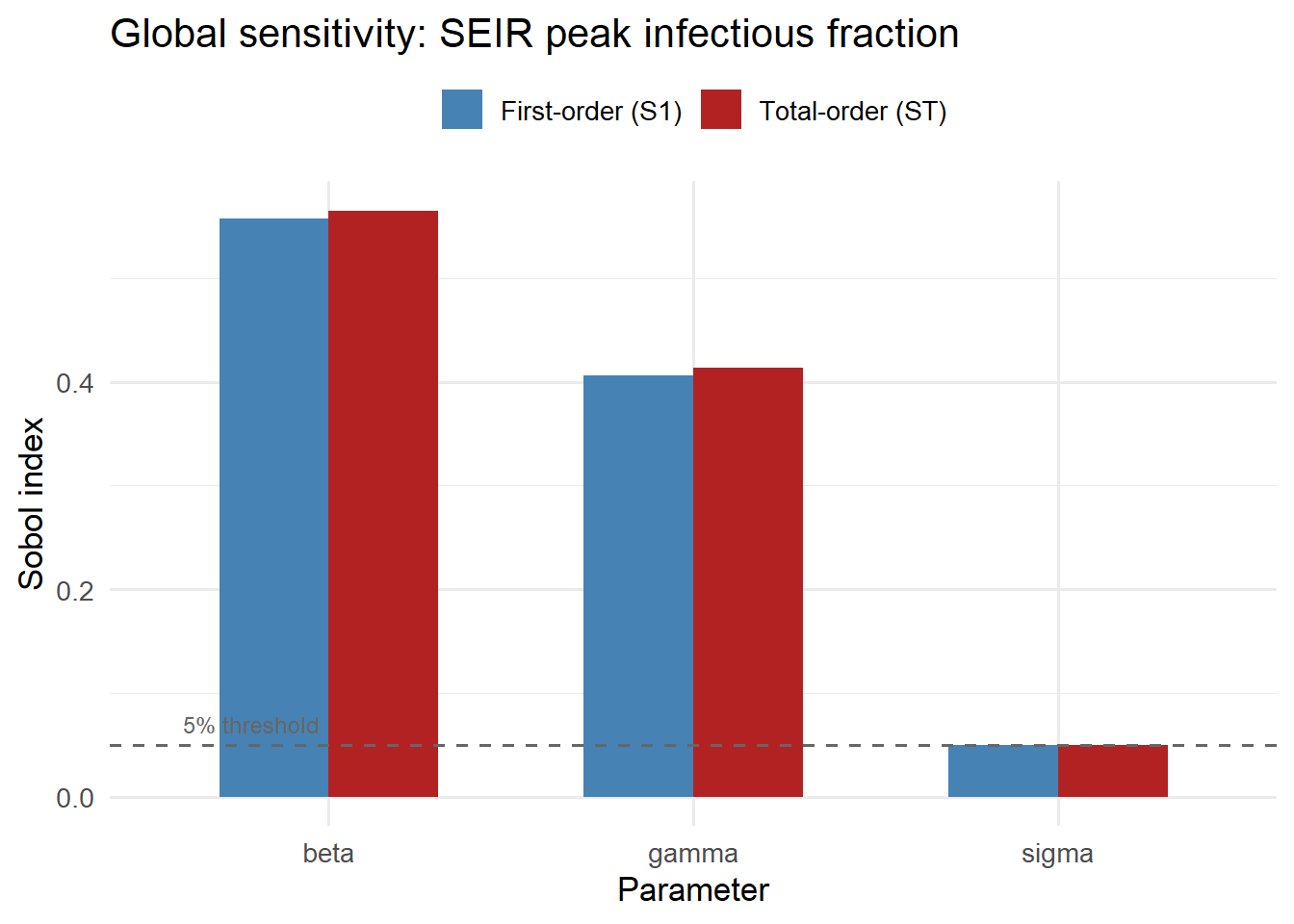
4 Morris screening for quick triage
When you have many parameters (20+), full Sobol estimation is expensive. The Morris method (2) is a cheap rank-order screen: run the model \(r(k+1)\) times (rather than \(2nk\)), and compute the mean \(\mu^*\) and standard deviation \(\sigma\) of elementary effects.
- High \(\mu^*\): parameter has large influence
- High \(\sigma\): parameter has nonlinear effects or interactions
morris_screen <- function(model_fn, ranges, r = 20, delta = 0.5) {
k <- length(ranges)
nms <- names(ranges)
EE <- matrix(0, nrow = r, ncol = k)
for (rep in seq_len(r)) {
# Random base point in [0, 1-delta]
x0 <- sapply(seq_len(k), function(i) runif(1, 0, 1 - delta))
# Evaluate at base
params0 <- mapply(function(x, r) r[1] + x * diff(r), x0, ranges)
y0 <- do.call(model_fn, as.list(setNames(params0, nms)))
# Randomly permute dimensions and apply +delta one at a time
perm <- sample(k)
x_current <- x0
for (j in perm) {
x_new <- x_current
x_new[j] <- x_current[j] + delta
params_new <- mapply(function(x, r) r[1] + x * diff(r),
x_new, ranges)
y_new <- do.call(model_fn, as.list(setNames(params_new, nms)))
EE[rep, j] <- (y_new - y0) / delta
y0 <- y_new
x_current <- x_new
}
}
data.frame(
parameter = nms,
mu_star = colMeans(abs(EE)),
sigma_ee = apply(EE, 2, sd)
)
}
morris_res <- morris_screen(seir_peak, ranges, r = 50)
ggplot(morris_res, aes(x = mu_star, y = sigma_ee,
label = parameter)) +
geom_point(size = 4, colour = "steelblue") +
geom_text(size = 4, nudge_x = 0.005, hjust = 0) +
labs(x = "μ* (mean |elementary effect|)",
y = "σ (SD of elementary effects)",
title = "Morris sensitivity screening: SEIR model") +
theme_minimal(base_size = 13)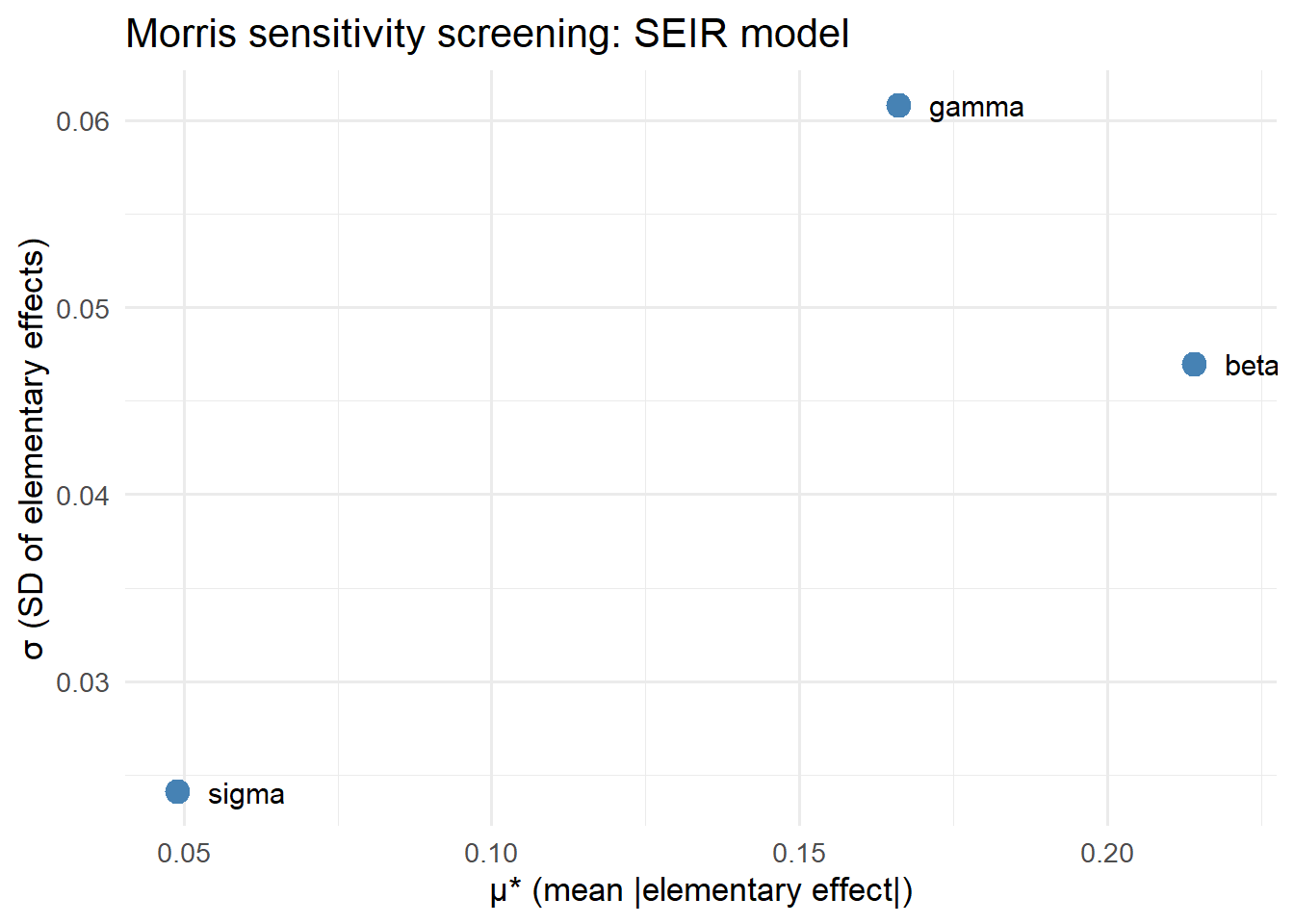
5 Packaging as a deliverable
A sensitivity analysis report for a client should include:
- Parameter ranges — what assumptions went into the SA
- Sobol indices table with confidence intervals (bootstrap)
- Ranking narrative — “transmission rate \(\beta\) explains 78% of forecast variance”
- Implications — which parameters are worth measuring better vs. fixing at nominal values
The sensitivity R package provides production-ready estimators with confidence intervals. The SALib Python library is the equivalent for Python-based models.